MOUSE ANTI-SARS-COV-2 SPIKE (S2) ANTIBODY (AD10)
Mouse anti SARS-CoV-2 Spike (S2) antibody (AD10) is a monoclonal antibody that recognizes the Spike subunit 2 (S2) glycoprotein of SARS-CoV-2, which is the causative agent of COVID-19. This antibody has been manufactured for use in ELISA immunoassay development and is suitable for use as a capture antibody in ELISA assays.
PRODUCT DETAILS – MOUSE ANTI-SARS-COV-2 SPIKE (S2) ANTIBODY (AD10)
- Antibody recognizes SARS-CoV-2 spike subunit 2 (S2) glycoprotein.
- Isotype – Mouse IgG1
- Immunogen was recombinant S2 protein expressed in HEK293 cells (REC31807, aa 685-1211).
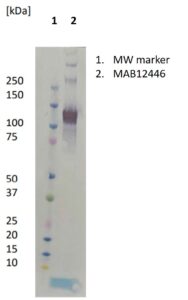
- Protein G purified from hybridoma culture supernatant.
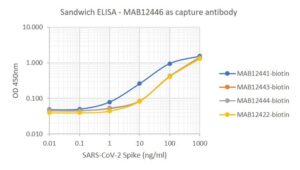
- Suitable for use as capture antibody in ELISA assay with MAB12422, MAB12441, MAB12443 or MAB12444 as the detection antibody.
BACKGROUND
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the causative agent of the coronavirus induced disease 19 (COVID-19) which emerged in China in late 2019, resulting in a worldwide epidemic (Zhou et al., 2020). SARS-CoV-2 is an enveloped positive-sense single-stranded RNA virus with a number of important structural proteins, including the envelope (E) protein, the membrane (M), the spike (S) protein, and the nucleoprotein (N). The S protein assists in the attachment of the virus to the human cell and comprises intracellular, transmembrane, and extracellular regions. The extracellular region contains the S1 receptor binding subunit (RBD) and the S2 membrane fusion subunit. Spike protein S2 mediates fusion of the virion and cellular membranes by acting as a class I viral fusion protein. Spike protein S2′ acts as a viral fusion peptide which is unmasked following S2 cleavage occurring upon virus endocytosis (Gui et al., 2017). Cytotoxic T-lymphocyte (CTL) epitopes reside within the S2 subunit of SARS-CoV and S2 may serve as an important antigen for inducing both humoral as well as cell-mediated immunity against SARS-CoV and SARS-CoV-2 (Poh et al., 2009; Ng et al., 2014). An immunogenic fragment in the S2 subunit of SARS-CoV has been used previously to generate mAbs which bind to the S protein through a novel epitope within the S2 subunit and had the ability to bind and cross-neutralize pseudotyped viruses expressing spike of human SARS-CoV, civet SARS-CoV and bat SL-CoV strains (Ng et al., 2014) and SARS-CoV-2 infected cells (Zheng et al., 2020). The Native Antigens monoclonal antibodies against SARS-CoV-2 subunit 2 (S2) have been manufactured for the development of improved diagnostic assays for COVID-19.
REFERENCES
- Gui M, Song W, Zhou H, et al. (2017). Cryo-electron microscopy structures of the SARS-CoV spike glycoprotein reveal a prerequisite conformational state for receptor binding. Cell Res. 2017;27.119–129.
- Ng OW, Keng CT, Leung CS, et al. Substitution at aspartic acid 1128 in the SARS coronavirus spike glycoprotein mediates escape from a S2 domaintargeting neutralizing monoclonal antibody. PLoS One. 2014;9(7):e102415.
- Ng O-W, Keng C-T, Leung CS-W, Peiris JSM, Poon LLM, Tan Y-J (2014) Substitution at Aspartic Acid 1128 in the SARS Coronavirus Spike Glycoprotein Mediates Escape from a S2 Domain-Targeting Neutralizing Monoclonal Antibody. PLoS ONE 9(7): e102415.
- Poh WP, Narasaraju T, Pereira NA, et al. Characterization of cytotoxic Tlymphocyte epitopes and immune responses to SARS coronavirus spike DNA vaccine expressing the RGD-integrin-binding motif. J Med Virol. 2009 Jul;81(7):1131-9.
- Zheng Z, Monteil VM, Maurer-Stroh S, Yew CW, Leong C, Mohd-Ismail NK, Cheyyatraivendran Arularasu S, Chow VTK, Lin RTP, Mirazimi A, Hong W, Tan YJ. Monoclonal antibodies for the S2 subunit (1):119–129.of spike of SARS-CoV-1 cross-react with the newly-emerged SARS-CoV-2. Euro Surveill. 2020 Jul;25(28):2000291.
- Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020 Mar;579(7798):270-273.